 |
NMR structure of a photoswitchable G-quadruplex |
2N9Q |
 |
Crystal Structure of Ephrin B4 (EphB4) Receptor Protein Kinase with Dasatinib |
6FNM |
 |
Crystal Structure of Ephrin B4 (EphB4) Receptor Protein Kinase |
6FNL |
 |
Crystal Structure of Ephrin B4 (EphB4) Receptor Protein Kinase with a pyrazolo[3,4-d]pyrimidine fragment of NVP-BHG712 |
6FNK |
 |
Crystal Structure of Ephrin B4 (EphB4) Receptor Protein Kinase with an isomer of NVP-BHG712 |
6FNJ |
 |
Crystal Structure of Ephrin B4 (EphB4) Receptor Protein Kinase with NVP-BHG712 |
6FNI |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with a pyrazolo[3,4-d]pyrimidine fragment of NVP-BHG712 |
6FNH |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with an isomer of NVP-BHG712 |
6FNG |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with NVP-BHG712 |
6FNF |
 |
Structural characterization of the Mycobacterium tuberculosis Protein Tyrosine Kinase A (PtkA) |
6F2X |
 |
Impact of IR active probes on PDZ3 and its ligand binding studied by NMR and X-ray crystallography |
5w72 |
 |
Crystal Structure of the third PDZ domain from the synaptic protein PSD-95 with incorporated Azidohomoalanine |
5MZ7 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 1g |
5NJZ |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 1j |
5NK0 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 1k |
5NK1 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2b |
5NK2 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 1l |
5NK3 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2c |
5NK4 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 1m |
5NK5 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2d |
5NK6 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2a |
5NK7 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2f |
5NK8 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2e |
5NK9 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2g |
5NKA |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 4a |
5NKB |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2h |
5NKC |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 2i |
5NKD |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 3a |
5NKE |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 3b |
5NKF |
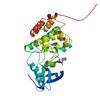 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 3d |
5NKG |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 3e |
5NKH |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with Compound 4b |
5NKI |
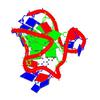 |
Solution structure of a human G-Quadruplex hybrid-2 form in complex with a Gold-ligand |
5MVB |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase |
5I9U |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with AGS |
5I9V |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with ANP |
5I9W |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with danusertib (PHA739358) |
5I9Z |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with dasatinib |
5I9Y |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with bosutinib (SKI-606) |
5I9X |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with alisertib (MLN8237) |
5IA0 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with MLN8054 |
5IA1 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with compound 66 |
5IA2 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with PD173955 |
5IA3 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with foretinib (XL880) |
5IA4 |
 |
Crystal Structure of Ephrin A2 (EphA2) Receptor Protein Kinase with golvatinib (E7050) |
5IA5 |
 |
NMR based solution structure of PTS system, galactitol-specific IIB component from methicillin resistant Staphylococcus aureus |
5GQS |
 |
Tetramerization domain of the Ciona intestinalis p53/p73-b transcription factor protein |
2MW4 |
 |
Two synonymous gene variants encode proteins with identical sequence, but different folding conformations |
4W9B |
 |
Crystal structure of Gamma-B Crystallin expressed in E. coli based on mRNA variant 2 |
4W9A |
 |
FXR with DM175 and NCoA-2 peptide |
4QE8 |
 |
FXR with CDCA and NCoA-2 peptide |
4QE6 |
 |
Crystal structure of mitochondrial NADH:ubiquinone oxidoreductase from Yarrowia lipolytica. |
4WZ7 |
 |
Crystal structure of SAH-bound Podospora anserina methyltransferase PaMTH1 |
4YMH |
 |
Crystal structure of SAM-bound Podospora anserina methyltransferase PaMTH1 |
4YMG |
 |
Apo-crystal structure of Podospora anserina methyltransferase PaMTH1 |
4QVK |
 |
Crystal structure of cAMP-dependent Protein Kinase A from Cricetulus griseus |
4WIH |
 |
Low resolution crystal structure of the FGFR2D2D3/FGF1/SSR128545 complex |
4J23 |
 |
Crystal structure of the electron transfer complex of cytochrome p450cam with putidaredoxin |
3W9C |
 |
The structure of the complex of cytochrome P450cam and its electron donor putidaredoxin |
2M56 |
 |
NMR solution structure of apo-MptpA |
2LUO |
 |
High resolution NMR solution structure of helix H1 of the human HAR1 RNA |
2LUB |
 |
High resolution NMR solution structure of helix H1 of the chimpanzee HAR1 RNA |
2LHP |
 |
Solution NMR Structure of Proteorhodopsin. |
2L6X |
 |
Crystal structure of guanine riboswitch C61U/G37A double mutant bound to thio-guanine |
3RKF |
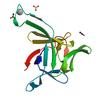 |
Structure of Interleukin 1B solved by SAD using an inserted Lanthanide Binding Tag |
3LTQ |
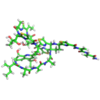 |
Thiostrepton, reduced at N-CA bond of residue 14 |
2L2Z |
 |
Thiostrepton, epimer form of residue 9 |
2L2Y |
 |
Thiostrepton, oxidized at CA-CB bond of residue 9 |
2L2X |
 |
Thiostrepton |
2L2W |
 |
Interleukin-1-beta LBT L3 Mutant |
3POK |
 |
NMR SOLUTION STRUCTURE OF A COMPLEX OF CALMODULIN WITH A BINDING PEPTIDE OF THE CA2+-PUMP |
1CFF |
 |
NMR SOLUTION STRUCTURE OF HEN LYSOZYME |
1E8L |
 |
GLU C180 -> ILE VARIANT QUINOL:FUMARATE REDUCTASE FROM WOLINELLA SUCCINOGENES |
2BS4 |
 |
GLU C180 -> GLN VARIANT QUINOL:FUMARATE REDUCTASE FROM WOLINELLA SUCCINOGENES |
2BS3 |
 |
Dual binding mode of pyridinylimidazole to MAP kinase p38 |
2EWA |
 |
QUINOL:FUMARATE REDUCTASE FROM WOLINELLA SUCCINOGENES |
2BS2 |
 |
Structure of ubiquitin solved by SAD using the Lanthanide-Binding Tag |
2OJR |
 |
Model for thiostrepton binding to the ribosomal L11-RNA |
2JQ7 |
 |
Ribosomal protein L11 from Thermotoga maritima |
2K3F |
 |
Human CDC37-HSP90 docking model based on NMR |
2K5B |
 |
HSP90 CO-CHAPERONE CDC37 |
2W0G |
 |
alpha-amylase inhibitor Parvulustat (Z-2685) from Streptomyces parvulus |
2KER |
 |
High resolution NMR solution structure of a complex of HIV-2 TAR RNA and a synthetic tripeptide in a 1:2 stoichiometry |
2KMJ |
 |
NMR solution structure of a 14-mer hairpin RNA with cUUCGg tetraloop |
2KOC |